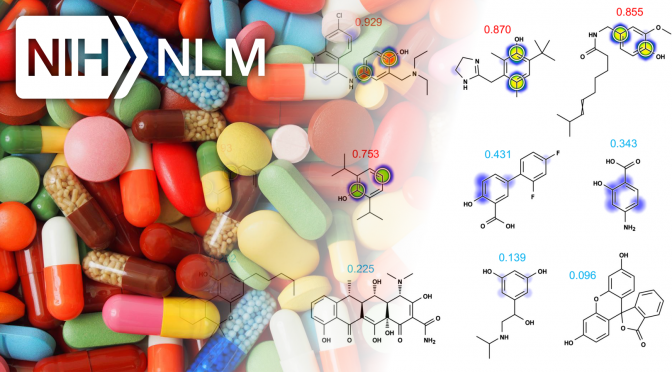
Metabolism and Reactivity (FUNDED)
Dr. Grover P. Miller Ph.D. and Dr. S. Joshua Swamidass M.D. Ph.D. are pleased to announce that our grant "Data and Tools for Modeling Metabolism and Reactivity" was funded�by the National Library of Medicine (NLM) of the NIH after receiving an outstanding�impact score of 21 (5th percentile) by the independent study section.
This … Continue Reading ››
ACS San Diego 2016
{{unknown}}
Marc Rubio and the Age-of-the-Earth Question
{{unknown}}
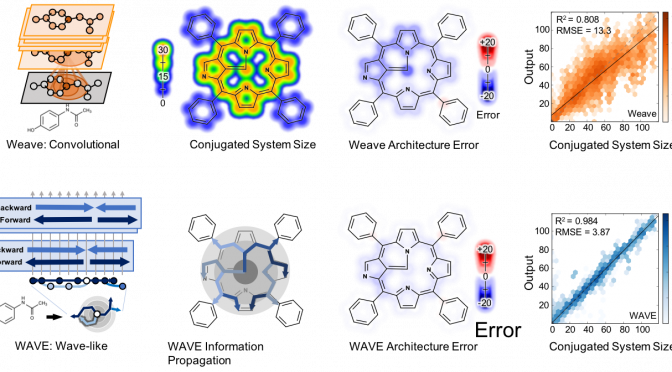
Update: Modeling Epoxidation of Drug-like Molecules with a Deep Machine Learning Network
This is a minor update�to figure 10 in a recently published paper:�Original Paper.
The revised figure�displays both the RMSE between each method and perfectly scaled prediction, and the R2 values for the Pearson correlation of the best-file line for that method. As measured by RMSE, the three methods have the same relative value as in … Continue Reading ››
Matthew Matlock Joins The Group
We are happy to have Matt Matlock officially join the group for his PhD. Matt has worked in the group over the last three years, but now that he is entering the PhD portion of his MD/PhD program, he will be with us full time.
Chemical Computing Group Excellence Award
{{unknown}}
Education: Initiatives to bridge faith and science
{{unknown}}
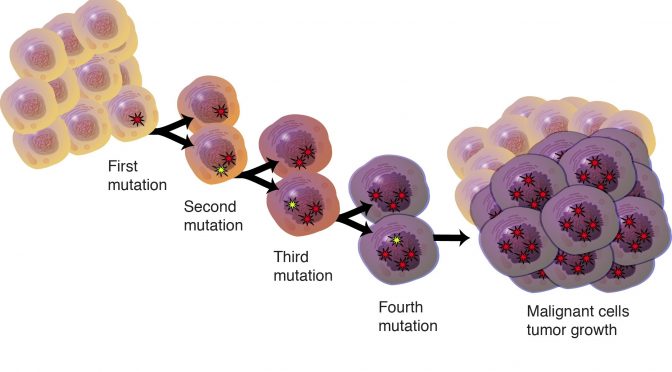
Statistically identifying tumor suppressors and oncogenes from pan-cancer genome-sequencing data.
{{unknown}}
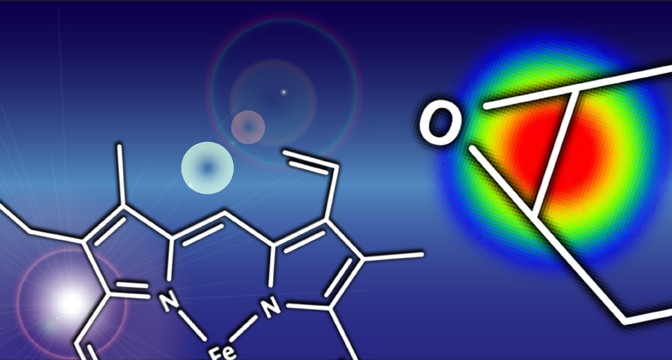
Modeling Epoxidation of Drug-like Molecules with a Deep Machine Learning Network
Citation:
Modeling Epoxidation of Drug-like Molecules with a Deep Machine Learning Network.�Tyler B. Hughes, Grover P. Miller, and S. Joshua Swamidass. �ACS Central Science.�DOI: 10.1021/acscentsci.5b00131
Abstract:�
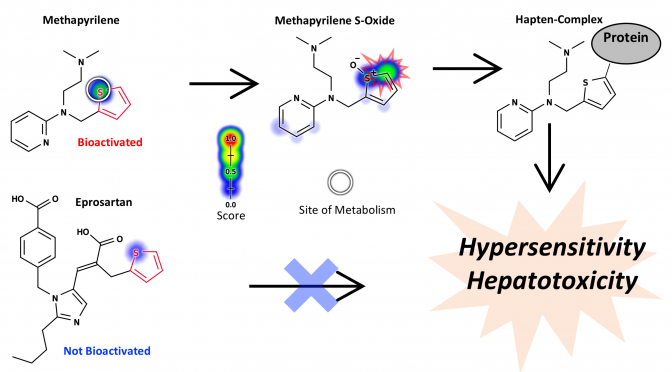
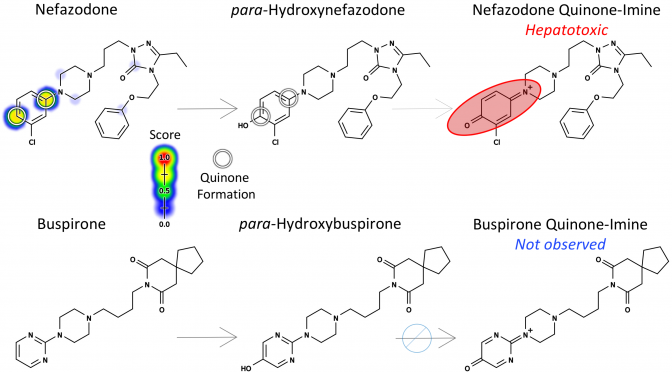
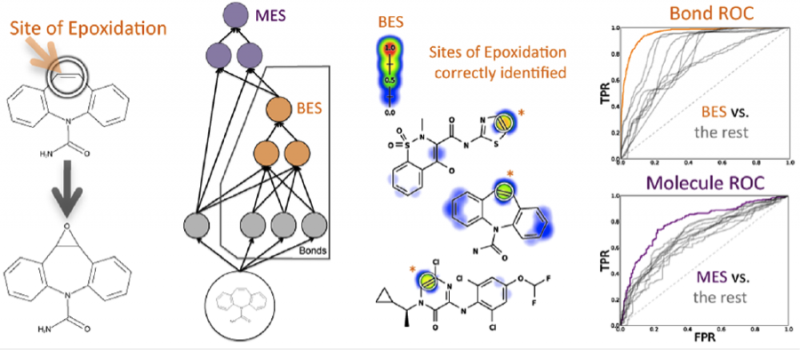 Drug toxicity is frequently caused by electrophilic reactive metabolites that covalently bind to proteins. Epoxides comprise … Continue Reading ››
Drug toxicity is frequently caused by electrophilic reactive metabolites that covalently bind to proteins. Epoxides comprise … Continue Reading ››
 Drug toxicity is frequently caused by electrophilic reactive metabolites that covalently bind to proteins. Epoxides comprise … Continue Reading ››
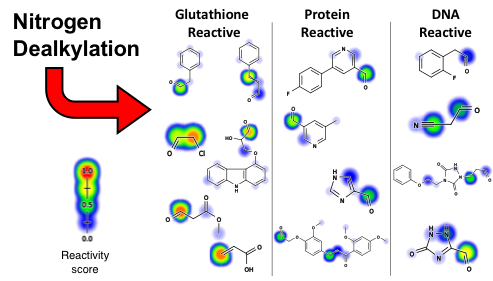
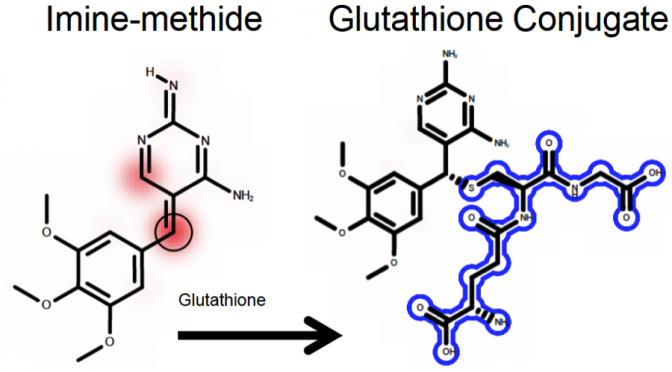
Drug toxicity is frequently caused by electrophilic reactive metabolites that covalently bind to proteins. Epoxides comprise … Continue Reading ›› Site of Reactivity Models Predict Molecular Reactivity of Diverse Chemicals with Glutathione
{{unknown}}